Building an atlas of all the cell types that make up the body typically requires multinational collaborations and massive budgets. But a technique that can analyse the genetic activity of hundreds of thousands of individual cells at a time has allowed one small team to produce a time-lapse atlas of an embryonic mouse’s cells over ten days of development. The atlas, which was created by three researchers in one year for approximately US$370,000, could help scientists to understand how stem cells turn into specific cell types, how organs develop and even how the body changes just after it is born.
The study, which was published in Nature in February1, “is impressive at many levels, both the scale of what they achieved and how they achieved it”, says Bertie Göttgens, a stem-cell biologist at the University of Cambridge, UK, who was not involved in the study.
Geneticist Jay Shendure at the University of Washington in Seattle doesn’t normally study mouse development. His laboratory is known for establishing molecular-biology techniques, including one called sci-RNA-seq3 that allows researchers to survey the assemblage of messenger RNA (mRNA), known collectively as the transcriptome, in individual cells.
Which single-cell analysis tool is best? Scientists offer advice
Instead of looking at whole cells, which would be difficult to keep intact through the process, scientists grind up a sample — in this case, a whole mouse embryo — and isolate its cell nuclei. They split these nuclei into individual dishes and add a different molecular tag to the mRNA in each dish. Next, they combine the nuclei, separate them again, mark each dish with a new tag and repeat. Eventually, each nucleus acquires a unique collection of tags — a molecular barcode — that the researchers can use to determine which tags define the cell’s transcriptome. They can then sequence these cells’ mRNA and construct a ‘tree’ that models how one cell type can turn into another, doing so across multiple animals of different ages on the basis of the genes they express.
Missing moments
Table of Contents
Two of Shendure’s lab members, postdoc Chengxiang Qiu and research scientist Beth Martin, decided to demonstrate sci-RNA-seq3 by charting the single-cell transcriptomes of embryonic mice during the animals’ roughly 19-day gestation period. At first, they collected embryos every 24 hours over a 5-day period, but the transcriptomes changed so much between time points that it was difficult to follow how stem cells turned into specific cell types over time2. Shendure likens it to a video that is missing too many frames: more like a stop-motion animation than a smooth progression.
So Martin and Qiu partnered with research scientist Ian Welsh at the Jackson Laboratory, a research institute and mouse-breeding facility in Bar Harbor, Maine. Welsh painstakingly collected 83 mouse embryos at 2–6-hour intervals over 10 days of gestation, from the point at which organs start to develop up until just after the animal’s birth. Welsh snap-froze the embryos and sent them to Seattle, where Martin collected single-cell transcriptomes. Qiu then mapped the data into trees that show when and how each of 190 cell types — liver or bone-marrow cells, for instance — originates in an embryo.

Smart software untangles gene regulation in cells
To flesh out the tree, the researchers integrated their data, which began eight days into gestation, with existing work from Shendure’s team and others that had mapped the transcriptomes of these and younger embryos. This added another 110,000 cells to the mix, and these data formed the tree’s ‘roots’, allowing the researchers to follow the branching of early stem cells into specific types seen in the older embryos.
The resulting atlas, containing the transcriptomes of mice across 45 time points, is now available for developmental biologists to study in more depth. With 12.4 million cells, it is the largest mouse-embryo atlas so far and is nearly one-quarter the size of the cell data collected by the Human Cell Atlas collaboration, which comprises 700 labs attempting to map all of the cells in the human body.
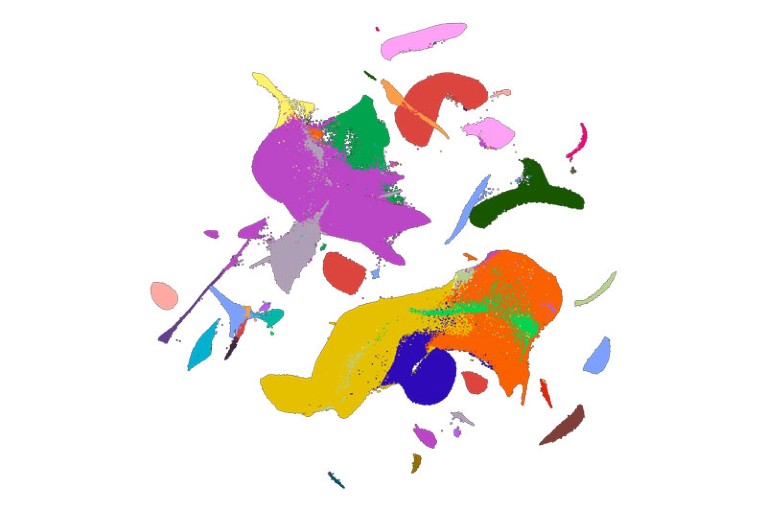

A 2D visualization of the mouse-atlas data set, with colours corresponding to 26 major cell types.Credit: C. Qiu et al./Nature
“It’s a fantastic resource for the community,” says cellular geneticist and Human Cell Atlas co-founder Sarah Teichmann at the Wellcome Sanger Institute in Hinxton, UK. Teichmann points out that there is still work to be done on the mouse atlas. Some time points have more complete transcriptomes than others, and the researchers have not yet separated mice by sex to look at those differences. But she says it will enable a number of studies, including the ability to compare mouse and human development. Shendure says he and his team plan to create single-cell atlases of juvenile and adult mice from conception to death.
Stress effects
Although Shendure and his group aim to let others conduct in-depth biological analyses of the data, they did note two phenomena in their paper. The point at which the transcriptome changed most dramatically, they found, was in the hour just after birth, which Shendure calls “the most stressful moment in your life”. Some of those differences were expected — lung and fat cells changed activity to cope with being outside the uterus, for instance — but other changes are still unclear.

How single-cell multi-omics builds relationships
Pure luck led them to another finding. To get the timing just right, Welsh typically delivered the mice by caesarean section. But one day, he returned from lunch to an unexpected nest of newborn pups. Martin processed the mice anyway and found that their transcriptomes were significantly different from those of mice born by caesarean section. Those differences could explain the variation in health outcomes seen between people who were born by these two methods, the researchers say.
Yonatan Stelzer, an epigeneticist at the Weizmann Institute of Science in Rehovot, Israel, says the study is encouraging for future efforts to map the cells of individual organs or tissues. The next step for embryos, he says, will involve not only studying how cells develop over time, but also following them through space in 3D, tracking how they split and move to form a whole mouse. Future research, he adds, could also investigate questions such as how two cells with similar transcriptomes end up with different fates to become the right or left eye, for instance. “We’re still far from solving the entire embryonic puzzle,” he says.
