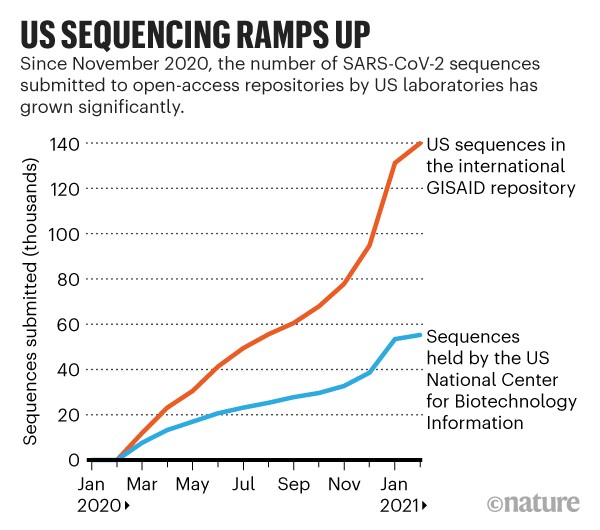
For the scientists who have spent the past year poring over hundreds of thousands of coronavirus genomes, the United States has been an enigma. Despite having world-leading genome sequencing infrastructure and experiencing more COVID infections than any other country, the United States has until recently lagged far behind in sequencing coronavirus genomes and spotting worrisome variants.
But in recent weeks, US researchers have identified a host of new variants, including in California, New York State, Louisiana and elsewhere. And they are continuing to ramp up SARS-CoV-2 sequencing efforts.
That has brought another challenge: making sense of the variants that are discovered. They carry potentially worrying mutations and may be becoming more common, but a dearth of data on how the variants are spreading means the threat they pose is unclear.
“It’s a Wild West,” says Jeremy Kamil, a virologist at Louisiana State University in Shreveport who co-led a team that, last month, spotted a fast-rising variant in Louisiana, New Mexico and elsewhere. In the absence of clear data on a variant’s behaviour, “it’s as if there’s an unofficial policy that every variant is a variant of concern until proven otherwise”, says Kamil.
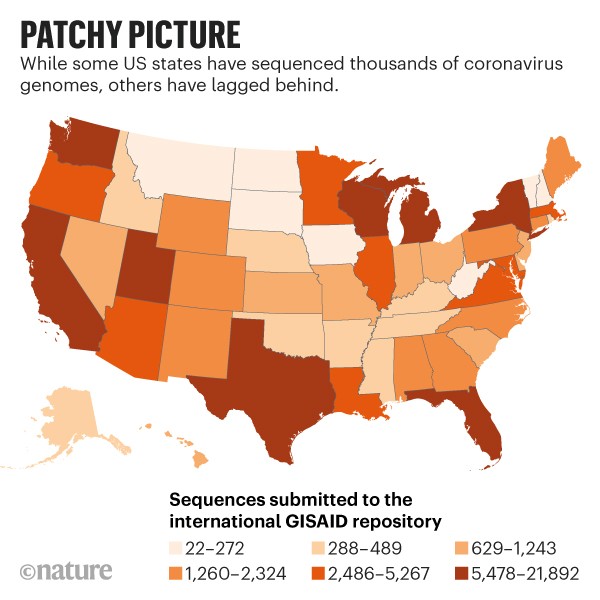
Part of the challenge is the decentralized nature of US coronavirus sequencing and surveillance efforts. “Right now it’s individual labs or states or cities doing their part,” says David Ho, a virologist at Columbia University in New York City, whose team last week identified a variant in the city with a mutation that could compromise immune responses. As a result of this piecemeal effort, states such as New York, California and Washington have contributed thousands of sequences each, while others such as Iowa, Tennessee and New Hampshire have obtained sequences from far fewer of their COVID cases.
Ho and other US researchers look enviously at the United Kingdom, where a nationwide sequencing effort — which works closely with public-health, medical and research institutions to make sense of variants — has generated more than 300,000 coronavirus genomes. Thanks to the fine granularity of its data, the UK effort showed in late 2020 that the variant called B.1.1.7 clearly spread faster than the circulating strains it would go on to displace; subsequent research has suggested that B.1.1.7 might be deadlier but does not compromise vaccines.
“I don’t think we have anything like that,” says Ho. He hopes that the US National Institutes of Health and the Centers for Disease Control and Prevention (CDC), the federal agencies responsible for biomedical research and public health, “will get the country moving in a more concerted fashion”. A CDC-led effort launched in November is aiming to sequence around 7,000 samples per week — a goal met for the first time in late February — and eventually 25,000.
Worrying mutations
Table of Contents
In the absence of clear-cut epidemiological or medical data, scientists can gauge some of a variant’s potential threat by the mutations it carries. Researchers have drawn up a growing list of mutations that might boost transmission or help a virus evade immune responses, based on laboratory and epidemiological studies.
The variant that Ho’s team identified in New York, also known as B.1.526, carries a notorious mutation called E484K that has been found in variants identified in South Africa and Brazil. Studies by multiple labs have shown that the E484K change — which is in a portion of the coronavirus spike protein that recognizes host cells — weakens the potency of antibodies that can ordinarily disable the virus. That could help explain observations that similar variants in South Africa and Brazil are behind cases of reinfection and reduced vaccine efficacy in field trials.
On the basis of those concerns, a team led by Ho and Columbia University microbiologist Anne-Catrin Uhlemann set up a surveillance network to identify viruses carrying E484K in New York City. The first cases of the B.1.526 variant appeared in November, growing to 5% of the city’s total cases in mid-January and 12% by February. In public sequencing databases, the researchers found B.1.526 up and down the northeast coast of the US and as far away as Singapore. The notoriety of E484K also inspired a team led by Pamela Bjorkman and Anthony West, structural biologists at the California Institute of Technology in Pasadena, to trawl through public sequencing data, where they spotted the lineage emerging in New York.
Ho concedes that B.1.526 needs much more study. It has not yet been shown to dodge immune responses and its apparent rise in frequency may be unrelated to any biological property. “It’s taken months for the UK variant [B.1.1.7] to be shown to be more transmissible and more virulent. I think we would need to do the same,” he says.
Rarer variants
Beefed-up US sequencing efforts are turning up variants with new or rarely seen mutations that are harder to make sense of. Kamil teamed up with researchers in New Mexico because they had also observed rising numbers of cases caused by a variant, which they dubbed Pelican, bearing a mutation they hadn’t seen before. They identified several other variants in US sequencing data that carried a similar change. (All were given bird nicknames, including Robin, Yellowhammer and Mockingbird.)
The mutation, called Q677P, sits near a region of the spike protein that needs to snap in two to allow the viral particle to enter a host cell. Mutations in this region occur in fast-spreading variants such as B.1.1.7, but Kamil says the Pelican variant is, for now, one to watch rather than worry over. “It’s too early to say with any scientific confidence that it’s a particularly worrisome mutation,” he says.
Last week, researchers in California raised a red flag over variants found there that carry a spike protein mutation called L452R1. A team at the University of California, San Francisco (UCSF) found that a variant with the mutation was rising rapidly in one city neighbourhood, from being present in 16% of sequenced samples in November to more than half in mid-January. Another UCSF team found, in lab tests, that a variant with the L452R mutation was more infectious and less susceptible to antibodies, according to media reports.
But many researchers have expressed scepticism of the significance of the L452R variants. The mutation hasn’t turned up in lab studies that have flagged several other worrisome changes, such as E484K, and the same L452R mutation has popped up elsewhere in the United States and not grown rapidly, says Jeffrey Barrett, a statistical geneticist at the Wellcome Sanger Institute in Hinxton, UK. “It’s probably not going to be one of these ones that is fundamentally problematic,” he says. “It’s a question of waiting and watching to see what happens in California and elsewhere.”
While researchers try to make sense of newly discovered US variants, redoubled sequencing efforts are also turning up more cases linked to global variants of concern. So far, researchers in the US have uncovered only a handful of cases linked to immune-evading variants identified in South Africa and Brazil. But cases of the B.1.1.7 variant identified in the United Kingdom are rising steadily — a pattern repeated in other countries in Europe and the Middle East.
Which of these variants will predominate is anyone’s guess, say researchers, but as vaccination ramps up, susceptible variants such as B.1.1.7 could dwindle, while those that can partly evade immunity might spark regional flare-ups. “I don’t think we’re going to have years of ‘New York variants’ and ‘California variants’, says Barrett. Figuring out what is happening will depend not only on sequencing more samples, but also on building the capacity to understand them. “The US must do a better job along these lines,” says Ho.