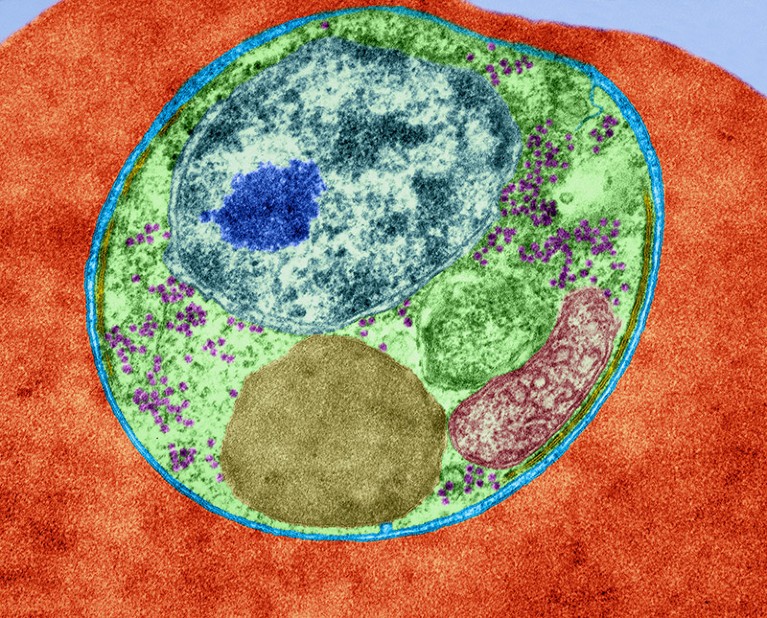
The malaria parasite Plasmodium falciparum infecting a red blood cell.Credit: Dennis Kunkel Microscopy/Science Photo Library
Researchers have sequenced the mitochondrial genome of the deadliest form of malaria from an ancient Roman skeleton. They say the results could help to untangle the history of the disease in Europe.
It’s difficult to find signs of malaria in ancient human remains, and DNA from the malaria-causing parasite Plasmodium rarely shows up in them. As a result, there had never been a complete genomic sequence of the deadliest species, Plasmodium falciparum, from before the twentieth century — until now. “P. falciparum was eliminated in Europe a half century ago, and genetic data from European parasites — ancient or recent — has been an elusive piece in the puzzle of understanding how humans have moved parasites around the globe,” says Daniel Neafsey, who studies the genomics of malaria parasites and mosquito vectors at the Harvard T.H. Chan School of Public Health in Boston, Massachusetts.
Malaria has long been a leading cause of human deaths. “With the development of treatments such as quinine in the last hundreds of years, it seems clear [humans and malaria] are co-evolving,” says Carles Lalueza Fox, a palaeogenomicist at the Institute of Evolutionary Biology in Barcelona, Spain. “Discovering the genomes of the ancient, pre-quinine plasmodia will likely reveal information about how they have adapted to the different anti-malarial drugs.”
Ancient pathogen
Table of Contents
There are five malaria-causing species of Plasmodium, which are thought to have arisen in Africa between 50,000 and 60,000 years ago, and then spread worldwide. Most researchers agree that they reached Europe at least 2,000 years ago, by the time of the Roman Empire.
Plasmodium falciparum “has significantly impacted human history and evolution”, says Neafsey. “So, that makes it particularly important to discover how long different societies have had to deal with [it], and how human migration and trade activities spread it.”
Researchers can glean valuable information about the origin, evolution and virulence of the parasite from DNA extracted from the ancient remains of infected people. But it is difficult to know where to look: it is not always obvious whether a person was infected with Plasmodium, and whether DNA can be recovered depends on how well it has been preserved.
In a preprint posted on the server bioRxiv1, a team of researchers led by a group at the University of Vienna identified the first complete mitochondrial genome sequence of P. falciparum from the bones of a Roman who lived in Italy in the second century ad, known as Velia-186.
Plasmodium falciparum had been detected in Velia-186 in a previous study2. The authors of the latest preprint extracted the parasite’s DNA from the body’s teeth, and were able to identify 5,458 pieces of unique genetic information that they combined to get a sequence covering 99.1% of the mitochondrial genome. They also used software to compare the genome with modern samples, and found that the Velia-186 sequence is closely related to a group of present-day strains found in India.
Carried by migration
The researchers say their findings support a hypothesis that P. falciparum spread to Europe from Asia around at least 2,000 years ago3. The Indian strains “were already present in Europe [then]; thus, a potential arrival with globalization episodes such as the Hellenistic period — when it is first described by Greeks — seems plausible”, says Lalueza Fox.
Neafsey says the work is a “technical tour de force” and an interesting addition to the limited field of ancient malaria genomics. But he adds that the results should be interpreted with caution because there are only a few samples, and points out that a genome sequence from DNA in the parasite’s cell nuclei, rather than its mitochondria, “might indicate a more complex story of parasite movement among ancient human populations”.
Lalueza Fox suggests exploring other potential sources of Plasmodium DNA, such as old bones, antique medical equipment and even mosquito specimens in museums. “The integration of genetic data from these heterogeneous sources will provide a nuanced view of this disease,” he says. “It would be interesting to see what lessons we can learn from the past on the strains and dispersals of this pathogen.”
