During embryonic development, new cell types emerge with stunning speed and robustness. The process of gastrulation — in which a single layer of cells gives rise to multiple ‘germ layers’ — is fundamental to the early development of most animals. Although studied for more than 150 years, many aspects of gastrulation remain elusive, not least a comprehensive understanding of the molecular factors governing the specification of the many cell lineages that emerge from this process. Writing in Cell, Mittnenzweig et al.1 densely sample gene expression in gastrulating mouse embryos over a 36-hour window and construct a continuous model of cell-lineage specification.
If we think of cells during gastrulation as characters in a silent film (and, indeed, there are beautiful films of gastrulation2), how can we understand the internal monologues and ever-changing motivations of the personas on the screen? Only in the past five years or so, with the emergence of technologies that characterize the molecular profiles of individual cells, have we been in a position to fully monitor cells’ ‘inner lives’ throughout gastrulation, as cell lineages develop. One such technology is single-cell RNA sequencing (scRNA-seq), which profiles the messenger RNA contents of individual cells.
Several key questions remain that could be addressed through single-cell techniques. For example, what is the precise timing of cell-type specifications in the developing embryo? Can we find a model that accurately describes these specifications? What are the principal molecular factors involved? And which of these factors ‘drive’ cell-type specifications, and which ‘respond’ to them?
In most animals, including mammals, three germ layers result from gastrulation: the ectoderm, the mesoderm and the endoderm. In the mouse, the pre-eminent model system for mammalian development, gastrulation begins about 6.5 days after fertilization (that is, embryonic day 6.5, or E6.5). Although we and others have performed scRNA-seq across early mouse development at what might seem like reasonable temporal resolution (for example, sampling every 6 hours from E6.5 to E8.53, or every 24 hours from E9.5 to E13.54), the pace of change during mouse development is so fast that this might be woefully inadequate. In our film analogy, this would be akin to watching a film but with only a handful of scenes narrated.
In this context, Mittnenzweig et al. set out to generate a continuous representation of cell-state dynamics during mouse gastrulation1. They applied scRNA-seq to 153 mouse embryos from E6.5 to E8.1, altogether profiling gene expression in about 33,000 individual cells. Because the accuracy of estimating embryo age on the basis of morphological landmarks is limited, the embryos’ ages were instead inferred from the molecular data, resulting in each embryo being assigned to one of 13 time points.
To increase the temporal resolution of their representation of development beyond a series of snapshots, Mittnenzweig et al. posited that cells within any given embryo are, to some extent at least, at different stages of developmental maturity relative to one another. The authors grouped cells according to their molecular similarity into 461 subsets termed ‘metacells’, each consisting of cells that were very similar, but that, notably, might have come from different embryos and/or from different time points. The authors then applied an algorithm to estimate the fraction of cells from each metacell at time t that ‘flow’ to other metacells at time t + 1. Crucially, the inferred flows between these metacells are continuous with respect to time, despite the temporally discrete nature of the embryos from which they were derived.
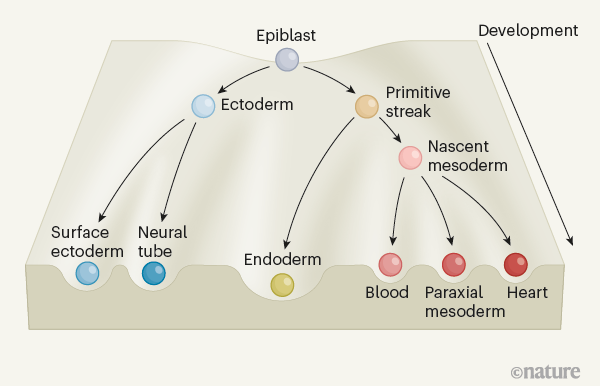
With this continuous model of mouse gastrulation in hand (Fig. 1), Mittnenzweig and colleagues are able to investigate several interesting questions. First, how and when do new cell types emerge during gastrulation, and what are the associated changes in the patterns of gene expression? For example, their model not only predicts that primitive erythroid cells (which give rise to early red blood cells) originate from a region called the primitive streak, but also constrains the timing of that contribution to before E6.7, and places in order the successive waves of expression of different transcription factors associated with this lineage.
Second, what are the characteristics of in vivo cell-type specification? Do new cell types emerge through a series of rapidly made ‘decisions’ between two different cell fates, resulting in the sharp, branch-like bifurcations that often appear in textbook flow charts of cell development, or are more-complex patterns observed, such as multifurcations and continuous transitions? Mittnenzweig et al. suggest the answer to be ‘all of the above’.
For example, the developmental trajectory of cells in the primitive streak bifurcates sharply, such that these cells become either mesodermal or endodermal cells (Fig. 1). By contrast, the differentiation of cells in the nascent mesoderm is inferred to be gradual and continuous, and with more than two destinations. The model also enables the inference of flows that change with time; for instance, before E7.1, epiblast cells overwhelmingly transition to acquiring primitive-streak fates, but shortly after that point, they mostly transition to acquiring ectodermal fates.
Finally, what are the molecular factors that underlie differentiation, and do individual factors act alone or in combination? The authors claim that, with the exception of some lineages (notably, the node, cardiomyocyte and haemato-endothelial lineages), the landscape of gastrulation is predominantly characterized by a dependence on overlapping combinations of factors, as well as on a gradual unfolding of commitment. For example, although cells of the nascent mesoderm progress into a spectrum of fates, these fates are not sharply separated from one another, and there is no clear delineation between the sets of transcription factors that seem to specify each fate. The authors propose that, rather than a series of specific factors governing a stepwise, hierarchical progression of specification, combinations of molecular factors regulate diverse mesodermal fates in a ‘fuzzy’ and almost probabilistic manner. To highlight the delicacy of this program, the authors carried out experiments in which inferred key regulators were genetically disrupted, which led to delayed differentiation of affected lineages.
Of course, all models have limitations, and this model has its own. First, its resolution is limited by the underlying data, although simply processing more embryos would address this. Second, its metacells and flows are inferred solely from the similarity of the transcriptional profiles of the cells, and so there is a risk of missing or misinterpreting certain bona fide relationships5. Particularly rapid changes in the nematode worm Caenorhabditis elegans, for instance, elude efforts to reconstruct lineages in ‘pseudotime’ — that is, ordering cells by their developmental stage rather than their age in real time6. Third, the model ignores cells’ spatial coordinates within embryos as well as their actual lineage relationships, two crucial aspects of development that are increasingly amenable to measuring and recording, respectively7,8.
Notwithstanding these limitations, the model of mouse gastrulation developed by Mittnenzweig et al. is impressive, and shows how continuous maps of complex differentiation landscapes might be recovered despite discrete sampling. Together with other work published in the past few years3,6,9, it represents a substantial step forward on the path to a complete understanding of cells’ inner lives during this most important of times in an animal’s life10.